Description
Pao is a LTR retrotransposon described in the genome of the domestic silkworm Bombyx mori and its name derives from the Chinese word for “running” (Xiong et al. 1993). Approximately 40 copies of Pao elements are present in the genome with most located outside the ribosomal DNA (rDNA) units (Xiong et al. 1993). Pao gives the name and belongs to Pao clade (Copeland et al. 2005) within Branch 2 of the Bel/Pao family (Llorens et al. 2009).
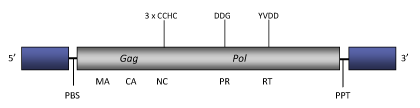
The genome of Pao is about of 4.8 Kb in size (4791 bp in lenght), including LTRs of 629 nt. The internal coding region displays one or two Primer Binding Site (PBS), complementary to a tRNATyr, a single 1158 amino acids ORF for gag and pol genes, and a Polypurine Tract (PPT) adjacent to the 3´LTR. The gag polyprotein presents three CCHC (Cysteine zinc finger array) motifs characteristic of the nucleocapsid domain, while pol shows absence of Integrase and part RNase H (Xiong et al. 1993).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
Mobile DNA II. (Craig et al. 2002)
|
|