Element:LLDAV
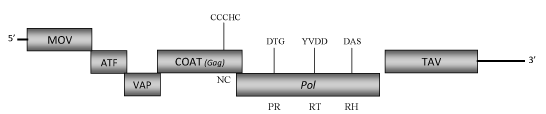
DescriptionLamium leaf distortion associated virus (LLDAV) is a plant pararetrovirus originally described in USA and detected consistently in the flowering plants "Purple dragon" (Lamium maculatum) in which the virus causes a growth disorder characterized by severe leaf deformation (Zhang et al. 2008). This pararetrovirus has been proposed to be a putative new member of Caulimovirus genus (Zhang et al. 2008) within Caulimoviridae family (International Committee on the Taxonomy of Viruses -ICTV- Fauquet et al. 2005). This is a perspective clearly supported by phylogenetic analyses inferred based on the COAT(GAG) and pol protein domains (Llorens et al 2009). The diseased plants show symptoms of severe leaf deformation, stunting and reduced vigor (Zhang et al. 2008). All attempts to transmit the virus by mechanical or graft inoculation or by aphid vector (Myzus persicae) have been failed as well as no viral integrated sequences have been identified into the plant host genome. Thus, the only way of LLDAV transmission seems to be by clonal propagation (Zhang et al. 2008). Morphologically, LLDAVs are spherical particles (45– 52 nm in diameter) composed of a genomic circular double-stranded DNA (7713 bp long). The genome encodes six Open reading frames (ORFs) whose size and organization are similar to those of known caulimoviruses (Zhang et al. 2008). The six ORFs potentially encode the putative movement protein (MOV, ORF I), the "aphid transmission factor" (ATF, ORF II), the "virion associated protein" (VAP, ORF III), the coat protein (COAT, ORF IV), the Pol polyprotein (Pol, ORF V) and the "Translational transactivator protein" (TAV, ORF VI) whose sequences show homologies with those of other Caulimoviruses (Zhang et al. 2008). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||