Ribonuclease H
Ribonuclease H (RNase H) is a hydrolytic enzyme widely distributed in both prokaryotes and eukaryotes (Johnson et al. 1986; Doolittle et al. 1989). In Ty3/Gypsy and Retroviridae and other LTR retroelements this enzyme is encoded as a part of the pol polyprotein and constitutes the C-terminal end of the Reverse Transcriptase (RT). RNase H is responsible for the hydrolysis of the original RNA template that is part of the RNA/DNA hybrid generated after the retrotranscription process in the viral life cycle.
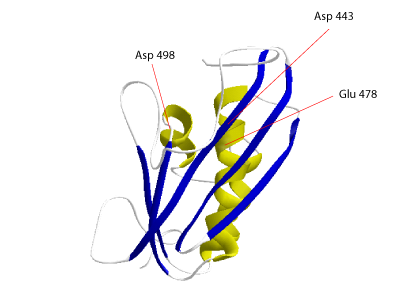
The three dimensional (3D) structure of the HIV-1 RNase H is characterized by four or five α-helices and five β-sheets that interact aligning in parallel to conform the active site (Davies et al. 1991). The activity of this enzyme normally requires the presence of divalent cations like Mg2+ or Mn2+ that bind to an active site constituted by a catalytic triad (Asp-443-Glu-478-Asp-498). These three residues have been proposed to be important in RNase H-mediated catalysis by HIV-1 RT (Mizarhi et al. 1990; Davies et al. 1991). Mutations in any of these resides inhibit the RNase H activity but have small effects on polymerase activity of the HIV-1 retrovirus ( Schatz et al. 1989; Mizarhi et al. 1990; Davies et al. 1991; Destefano et al. 1994).