Element:MiMV
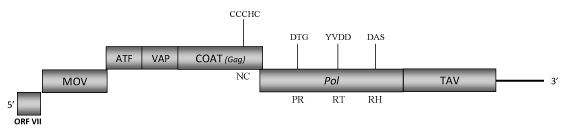
DescriptionMirabilis mosaic virus (MiMV) is a plant pararetrovirus isolated from the ornamental garden plants of Mirabilis species (family Nyctaginaceae) (Richins and Shepherd 1983). MiMV is member of the genus Caulimovirus, Caulimoviridae family (International Committee on the Taxonomy of Viruses -ICTV- Fauquet et al. 2005). Mirabilis infected plants display symptoms characterized by systemic mosaics, mottles, ringspots or necrosis. MiMV is transmitted mechanically as well as by aphid vectors (Myzus persicae) while it is not transmitted either by seed or by pollen (see also ICTVdb). Morphologically, MiMVs are spherical virions of approximately 50 nm in diameter (Brunt and Kitajima 1973) containing a genomic circular double-stranded DNA (7857 bp long). The genome exhibits four single-stranded discontinuities one in the (-) strand and the three others in the other (+) strand (Richins and Shepherd 1983; Dey and Maiti 1999) as well as seven ORFs (I, II, III, IV, V, VI, VII) (Dey and Maiti 1999). ORF I encodes a protein of 323 amino acids (aa) associated to cell-to-cell movement (MOV). ORFs II and III encode for two proteins of 165 and 124 aa, which respectively correspond with the "aphid transmission factor" (ATF) and the "virion associated protein" (VAP). ORF IV encodes the putative viral gag-like COAT protein and contains the typical caulimoviruses "RNA-binding" motif "C-X-C-X2-C-X4-H-X4-C" associated motifs (Llorens et al. 2009; Hull 1996; Bouhida et al. 1993).ORF V is a pol polyprotein of 674 aa displaying the aspartic protease (PR), reverse transcriptase (RT) and RNase H (RH) typical domains. ORF VI encodes a putative translational transactivator protein (TAV) of 536 aa long. ORF VII (nucleotides "7666-7857 and 1-36") encodes for a putative protein of unknown function (Dey and Maiti 1999). The intergenic region between TAV and ORF VII includes a promoter sequence whose expression activity has been demonstrated to be greater than that of the Cauliflower mosaic virus (CaMV) 35S promoter (Dey and Maiti 1999). StructureFigure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||