Element:Endovir1-1
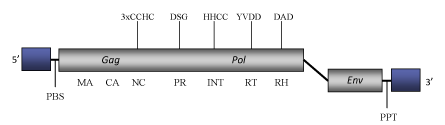
DescriptionEndovir1-1 is a LTR retroelement described in Arabidopsis thaliana (Peterson-Burch et al. 2000). Endovir1-1 belongs to the SIRE clade of Ty1/Copia plant LTR retrotransposons and potential retroviruses (Peterson-Burch et al. 2000; Fauquet et al. 2005;Peterson-Burch and Voytas, 2002), which is evolutionarily related to another clade called Oryco (Llorens et al. 2009). SIRE clade gives the name to a genus within the current ICTV classification of the Ty1/Copia into three genera – Pseudovirus, Hemivirus and Sirevirus (Boeke et al. 2005). The full-lenght genome of Endovir1-1 is 9093 bp in size, including two 549-bp LTRs (Peterson-Burch et al. 2000). The internal region of this element contains a Primer Binding Site (PBS) complementary to a tRNAiMet, a coding region and a Polypurine tract (PPT) upstream its 3' LTR. The coding region displays two ORFs. The first ORF encodes for the typical gag and pol domains of Ty1/Copia LTR retrotransposons, while the second ORF, separated from the first one by 1031 nucleotides, contains the env-like domains that exhibit a 24% amino acid identity to SIRE-1 other sirevirus element (Peterson-Burch et al. 2000). In comparison to other Ty1/Copia elements, a taxonomically relevant feature specific of Sire-like elements is their extremely large gag c-terminus consistent with the NC domain, which may display two or three CCHC arrays (Llorens et al. 2009). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||