Element:TF2
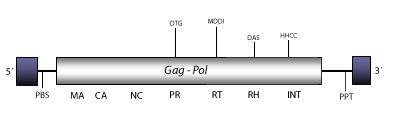
DescriptionTF2 is an active LTR retrotransposon characterized, along with TF1, in Schizosaccharomyces pombe (Levin, Weaver and Boeke 1990; Weaber et al. 1993). TF1 and TF2 are related to the Chromoviridae branch, term suggested (Marín and Lloréns 2000) to describe the Ty3/Gypsy elements bearing chromodomain-integrases (Malik and Eickbush 1999). The chromovirus branch is probably the most ancient phylogenetic pattern of Ty3/Gypsy retroelements (Gorinsek, Gubensek and Kordis 2004; 2005; Kordis 2005). The genomic structure of TF2 is 4.9 Kb in size, including LTRs of 349 nt. The internal region of this element displays a Primer Binding Site (PBS), a single Open Reading Frame (ORF) gag-pol, and a Polypurine Tract (PPT) adjacent to the 3´LTR (Levin, Weaver and Boeke 1990; Weaber et al. 1993). Neither, TF1 nor TF2 codify for chromodomain-containing integrases. The PBS of fungi and vertebrate chromoviruses, differs significantly from that used by plant chromoviruses. While plant chromoviruses use a methionine starting tRNA (iMet), fungi and vertebrate chromoviruses use their own self-priming mechanism to start the reverse transcription (Levin 1995 ; Butler et al. 2001). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||