Element:Tkm1
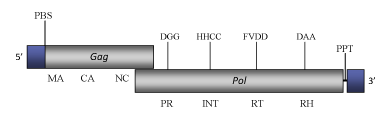
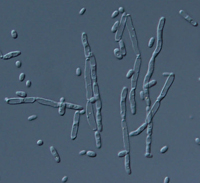
DescriptionTkm1 is a Ty1/Copia-like LTR retrotransposon, characterized in the genome of the yeast Kluyveromyces marxianus (the sexual form of Candida kefyr) (Neuvéglise et al. 2002), belonging to Ty (Pseudovirus) clade one of the three original genera on which the in-progrees Pseudoviridae classification is constituted (Boeke et al 2005). The genome of Tkm1 is about 6 Kb in size (5994 bp long), including two 385-bp LTRs. The internal region of this element presents a Primer Binding Site (PBS) complementary to a tRNAiMet, two overlapping Open Reading Frames (ORFs) and a final Polypurine Tract (PPT) (Neuvéglise et al. 2002). The first ORF, of 430 amino acids (aa), displays the gag domains, while the second larger ORF (1320 aa) presents the typical Ty1/Copia pol polyprotein structure: protease, integrase, reverse transcriptase and RNase H (Neuvéglise et al. 2002). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||