Description
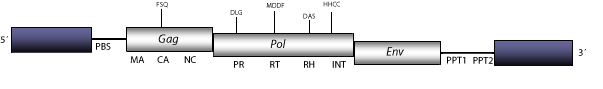
Cyclops is a LTR retroelement described in Pisum sativum (Chavanne et al 1998). Cyclops belongs to a lineage of possible retroviruses in plants (Wright and Voytas 2002), which is also related to other lineage (Tat) of plant LTR retrotransposons, providing evidence of a shared ancestral lineage originally described as class A by Wright and Voytas (1998). The figure below describes the Cyclops consensus. However, this elements has several subtypes; Subtype 2 is taken as representative. The genome of Cyclops-2 is 12.2 Kb in size, including LTRs of 1.5 Kb. Its internal region displays a Primer Binding Site (PBS) complementary to a tRNAAsp, three Open Reading Frames (ORFs) for gag, pol, and env genes, and two Polypurine Tracts, PPT1 and PPT2 adjacent to the 3´LTR (Chavanne et al 1998; Wright and Voytas 1998; 2002).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|