Element:rGmr1
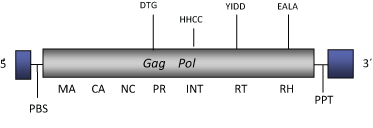
DescriptionrGmr1 is the counterpart of Gmr1 found in the genome of the zebrafish Danio rerio (Butler et al. 2001; Goodwin and Poulter 2002). These constitute an own clade, together with other elements from diverse vertebrates, possibly related via sequence similarity with Osvaldo clade (Butler et al. 2001; Goodwin and Poulter 2002; Llorens et al. 2009). rGmr1-like elements differ from other Ty3/Gypsy LTR retroelements in the fact that the integrase domain lies upstream of the reverse transcriptase domain, an arrangement which is characteristic of Ty1/Copia LTR retroelements (Butler et al. 2001; Goodwin and Poulter 2002). The genomic structure of rGmr1 is 4.8 Kb in size including LTRs of 21 nt. The internal region displays a Primer Binding Site (PBS), a single Open Reading Frame (ORF) that codes for a gag-pol polyprotein, and a Polypurine Tract (PPT) adjacent to the 3´LTRs. Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||