Element:Zeco2
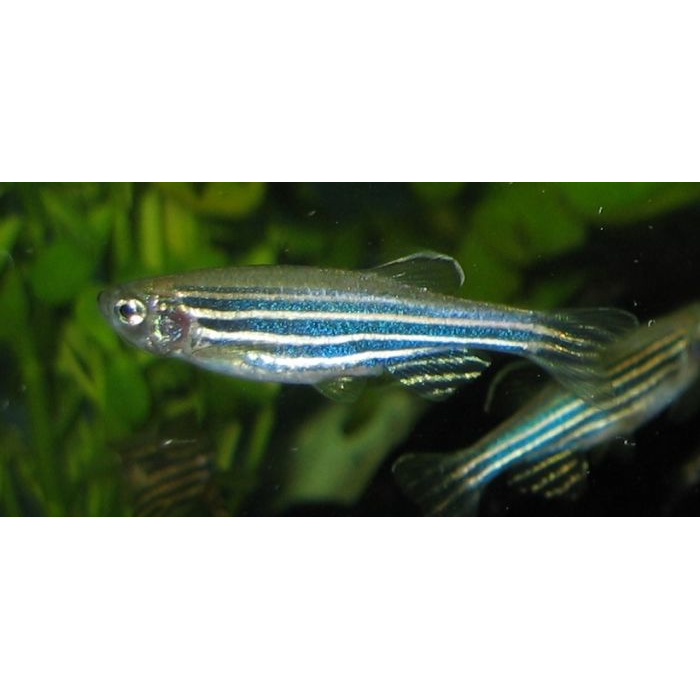
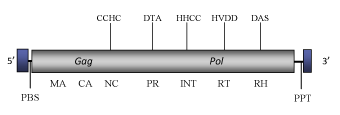
DescriptionZeco2 is a Ty1/Copia-like LTR retrotransposon described, together its closer related element Zeco1, in the genome of the zebrafish (Danio rerio) (Terrat et al. 2008). It belongs to GalEA clade a Ty1/Copia lineage specifically found in marin animals (Terrat et al. 2008). The genome of Zeco2 is about of 4.8 Kb in size (4840 bp long) including a 204-bp 5’ LTR and a 197-bp 3’LTR that flank an internal coding region. Although this region is disrupted by a several stop codons and frameshifts, it displays the typical gag and pol domains associated to Ty1/Copia LTR retrotransposons. A Primer Binding Site (PBS) complementary to a tRNAMet and a Polypurine Tract (PPT) are located respectively downstream and upstream to the 5’LTR and 3’LTR (Terrat et al. 2008). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||