Element:Tricopia
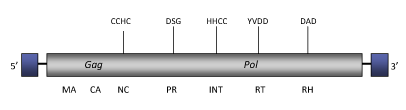
DescriptionTricopia is a LTR retrotransposon found in the genome of the red flour beetle Tribolium castaneum. It gives the name and constitutes a single clade within Branch 2 of the Ty1/Copia LTR retrotransposons family (Llorens et al. 2009). The genome of Tricopia is about of 4.5 Kb (4541 bp long), including LTRs of 260 (5’LTR) and 258 (3’LTR) bp long. The LTRs flanked internal region encodes for a single long ORF that, although is disrupted by some stop codons, contains the typical gag-pol domains structure of Ty1/Copia LTR retrotransposons (gag, protease, integrase, reverse transcriptase and ribonuclease H). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||