Description
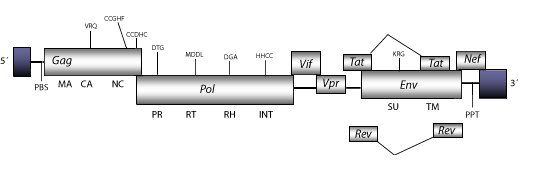
Simian Immunodeficiency Virus of Mandrill (SIVMND) is a complex retrovirus detected in the African mandrill (Tsujimoto et al. 1988). This retrovirus is similar to HIV-1 and other lentiviruses inductor of Acquired Immunodeficiency Syndrome (AIDS) in primates. The available genome of SIVMND is 9.2 Kb in size including LTRs of 692 nt (pro 5´LTR-R/U5 and pro 3´LTR-U3/R of 291 nt and 401 nt, respectively). The internal region of SIVMND displays a Primer Binding Site (PBS) complementary to a tRNALys, Open Reading Frames (ORFs) for the gag, pol and env genes characteristic of retroviruses, tat and rev (both expanded in two exons) and vif, which are three accessory genes common in other lentiviruses, vpr and nef accessory genes specifically observed in primate lentiviruses, and a Polypurine Tract (PPT) adjacent to the 3´LTR (Tsujimoto et al. 1989).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|