Element:SERV
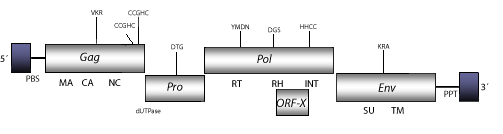
DescriptionSimian Endogenous Retrovirus of Mandrill (SERV) is a D-type retrovirus described from a mandrill genomic library (Van der Kuyl et al. 1997). Later PCR analyses also revealed sequences related to SERV in Old World monkeys of the Cercopithecinae subfamily. This retrovirus belongs to the genus Betaretrovirus, which comprises both B- and D-type retroviruses. Interestingly, D-type betaretroviruses present a common surface receptor also used by the baboon and cat endogenous C-type gammaretroviruses (BaEVM, RD114), cellular infection by any of them can interfere with other D-type members, and also with BaEVM and RD114 (Chatterjee and Hunter 1980; Sommerfelt and Weiss 1990). This evidence seems to be related with the high similarity displayed between env polyproteins encoded by gammaretroviruses and D-type betaretroviruses, indicating that betaretroviruses and gammaretroviruses may have been involved in an ancient recombinatorial event (Sonigo et al. 1986; Fauquet et al. 2005) or that they share a common ancestor (see env tree in section "phylogenies"). The genomic structure of SERV is 8.4 Kb size, including LTRs of 346 nt. The internal region displays a Primer Binding Site (PBS) complementary to a tRNALys3; four Open Reading Frames (ORFs) for gag, dut/pro, pol, and env genes, and a Polypurine Tract (PPT), as well, adjacent to the 3′LTR (Van der Kuyl et al. 1997; Elder et al. 1992; Payne and Elder 2001). We also have found that SERV also presents a putative accessory gene Orf-x that is common among many other betaretroviruses, and that was originally described in JSRV betaretroviruses (Bai et al. 1999; Rosati et al. 2000). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||