Description
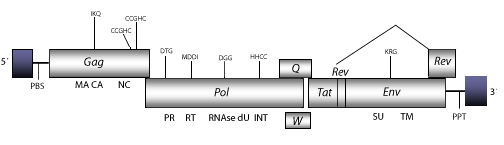
Ovine Maedi Visna Virus (SA-OMVV) is a sheep lentivirus found in South Africa and similar to Visna viruses of sheep. This genomic structure of this retrovirus is 9.2 Kb in size including LTRs of 499 nt (pro-5´LTR-R/U5 and pro3´LTR-U3/R of 152 nt and 347 nt, respectively. The internal region displays a Primer Binding Site (PBS) complementary to a tRNALys1,2, Open Reading Frames (ORFs) for gag, pol and env genes characteristic of retroviruses, tat and rev and orfq (vif or sor), which are three accessory genes common in other lentiviruses,an orfw also displayed but corrupted in other sheep lentiviruses, and a Polypurine Tract (PPT) adjacent to the 3′LTR (Saltarelli et al. 1990; Querat et al. 1990). As other non-primate lentiviruses, this retrovirus also displays between the RNase H and the INT an ORF coding for a dUTPase (dU) similar to that observed in betaretroviruses, N-terminal to the PR domain (Elder et al. 1992; Payne and Elder 2001 and references therein).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|