Element:PCSV
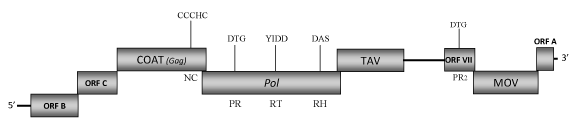
DescriptionPeanut chlorotic streak caulimovirus (PCSV or PCISV) is a plant pararetrovirus isolated from groundnut (Arachis hypogaea) (Reddy et al. 1993). PCSV is member of the genus Soymovirus within the Caulimoviridae family (International Committee on the Taxonomy of Viruses -ICTV- Fauquet et al. 2005). It is an economically important virus in tropical countries because it not only infects peanut, but also a broad host range of plants. This includes beans (Phaseolus vulgaris), cowpeas (Vigna unguiculata), soybean (Glycin max), Spinach (Spinacia oleracea), Datura stramonium, Petunia hybrid, and several Nicotiana species in which PCSV may provoke different symptoms such as prominent vein clearing, blistering, distortion and mottling, veinal necrosis (Reddy et al. 1993). PCSV can be transmitted by mechanical means to several Leguminosae and Solanaceae plants (Reddy et al. 1993). Morphologically, PCSV constitutes spherical virus particles 52-53 nm in diameter composed of a genomic 8174 bp dsDNA that contains two single-stranded discontinuities (Reddy et al. 1993). Its genome is organized in eight ORFs (I, A, B, C, IV, V, VI, VII). ORF I (MOV) encodes for a 320 amino acids (aa) cell-to-cell movement protein (Mushegian et al, 1995; Ducasse et al. 1995); ORFs A, B and C potentially encode for three proteins whose functions remain still unclear. Studies realized with infectious PCSV mutants showed that ORF A and C are essential for some aspects of virus life cycle, while ORF B is dispensable for infection (Mushegian et al, 1995). As to date an insect vector for PCSV is still unknown, it has been speculated with the possibility that ORF B might play a role in insect transmission (this remains to be determined). The coat protein gene (ORF IV) is one of the more conserved genes among caulimoviruses, it contains the characteristic plant pararetroviral "RNA-binding" motif (Hull 1996; Bouhida et al. 1993; Llorens et al. 2009). The predicted product of ORF V is a pol polyprotein displaying the aspartic protease (PR), reverse transcriptase (RT) and RNase H (RH) domains. ORF VI encodes for a putative translational transactivator protein (TAV) of 420 aa (Maiti et al. 1998). ORF VII encodes a protein of 143 aa with unknown function although our analyses predict that ORF VII may be an aspartic protease additional to that observed in the Pol gene. Interestingly, a similar feature has also been observed in other soymoviruses such as Soybean chlorotic mottle virus (SbCMV) and Blueberry red ringspot virus (BBRV) and in the tungrovirus Rice tungro bacilliform virus (RTBV). It is not clear however if these sequences are functionally active proteases or play other roles. Upon this, a previous study demonstrates that deletion of ORF VII in infectious PCSV mutants is correlated with the increased symptom severity (Mushegian et al, 1995). StructureFigure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||