Description
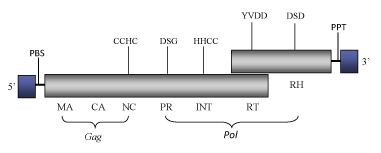
Osser is the first complete algae Ty1/Copia-like retroelement described in the green alga Volvox carteri (Lindauer et al. 1993). Osser constitutes a one-single-sequence clade within Branch 2 of the Ty1/Copia LTR retrotransposons family (Llorens et al. 2009) as it has no phylogenetic relatives to date reported. The full-length Osser genome is 4885 bp in size, including two identical 197-bp LTRs. The internal region of this element presents a Primer Binding Site (PBS) for a tRNAMet, two overlapping Open Reading Frames (ORFs) containing the typical gag and pol domains of Ty1/Copia LTR retrotransposons: gag (in which is present the zinc finger motif of Cys-His type characteristic of the nucleocapside domain), protease, integrase, reverse transcriptase and ribonuclease H; and a Polypurine Tract (PPT) upstream its 3’ LTR (Lindauer et al. 1993).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|