Element:Koco
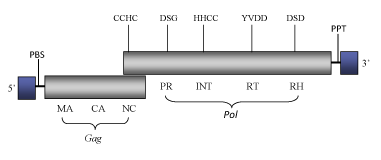
DescriptionKoco (koepferae Copia) is a Ty1/Copia LTR retrotransposon found in the genome of Drosophila koepferae (Francino and Cabre, Unpublished data, GenBank accession X96971). This element belongs to Copia clade, which gives the name to a genus within the current ICTV classification of the Ty1/Copia into three genera – Pseudovirus, Hemivirus and Sirevirus (Boeke et al. 2005). The genome of Koco is 4982 bp in size, including LTRs of 248 (5’LTR) and 247 nts long (3’LTR). This element contains a Primer Binding Site (PBS) for a tRNAMet, two overlapping Open Reading Frames (ORFs) containing the typical gag and pol domains of Ty1/Copia LTR retrotransposons: gag, protease, integrase, reverse transcriptase and ribonuclease H; and a Polypurine Tract (PPT) upstream its 3’ LTR (see also, Llorens et al. 2009). Structure
Related literature |
|
|||||||||||||||||||||||||||||||||||||||||