Description
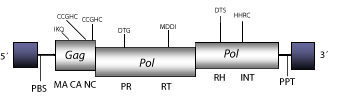
Gulliver is a LTR retrotransposon characterized in the genome of Schistosoma japonicum (Laha et al. 2001). Gulliver belongs, along with other LTR retrotransposons, to Mag a large Ty3/Gypsy cluster widely distributed in both protostomes and deuterostomes (for more details, see Malik and Eickbush 1999; Volff et al. 2001; Tubio, Naveira and Costas 2004). The genomic structure of the full-length genome of Gulliver is 4.8 Kb in size including LTRs of 259 nt. The internal region of this element displays a Primer Binding Site (PBS) complementary to a tRNA Met, two Open Reading Frames (ORFs) for gag, and pol genes, and a Polypurine Tract (PPT) adjacent to the 3´LTR, each LTR includes promotor sequences for the RNA polymerase II, a CAAT signal, and a TATA box (Laha et al. 2001).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|