Description
CoDi7.1 is a Ty1/Copia LTR retrotransposon described in the genome of the marine pinnate diatom Phaeodactylum tricornutum (Maumus et al. 2009; Bowler et al. 2008). It belongs to CoDi-I clade (Maumus et al. 2009), within Branch 1 of the Ty1/Copia family, that represents an interesting group of elements carrying a putative chromodomain at the C-terminus of the pol-INT domain (Llorens et al. 2009), similar to those coded by almost all Ty3/Gypsy chromoviruses which probably use this feature for chromatin integration (Gao et al. 2008).
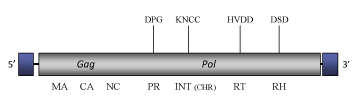
The genome of CoDi7.1, of about 6 kb in size (5954 bp long), includes 257-bp LTRs bounding a single long Open Reading Frame (ORF) of 1789 amino acids that contains the gag and pol typical structure of the Ty1/Copia LTR retrotransposons (gag, protease, integrase, reverse transcriptase and ribonuclease H, for more details, see Maumus et al. 2009).
Structure
Figure not to scale. If present, long terminal repeats (LTRs) have been highlighted in blue. Amino acid motifs noted with lines indicate the conserved residues in each protein domain, abbreviations below mean:
| MA=matrix
|
PR=protease
|
DU or DUT=dUTPase
|
TM=transmembrane
|
TAV or IBMP=transactivator/viroplasmin or inclusion body matrix protein
|
| CA=capsid
|
RT=reverse transcriptase
|
INT=Integrase
|
CHR=chromodomain
|
| NC=nucleocapsid
|
RH=RNaseH
|
SU=surface
|
MOV=movement protein
|
| PPT=polypurine tract
|
PBS=primer binding site
|
ATF=aphid transmission factor
|
VAP=virion associated protein
|
Related literature
|
|